The cost of sequencing has dropped precipitously over the last five years, the cost of library prep has not moved by anywhere near as much. For some experiments this is a major headache, particularly in small genomes, metagenomics, and for us, RNA-seq. Reducing the cost of consumables in kits is one way to bring down prices, but the cost savings get less and less if the protocol requires a person to spend a week in the lab; developing novel methods that are simpler is likely to have more impact. In Simultaneous generation of many RNA-seq libraries in a single reaction by Shishkin et al (Nat Methods 2015), a simple additional RNA-ligation step means 100's of samples can be processed in a one tube stranded RNA-seq library prep.
RNAtag-seq explained: Most of us are making RNA-seq libaries using one of the strand-marking dUTP methods, where each library is taken through the steps of library prep in Eppendorf's or 96-well plates. Scientists at the Broad Institute and CalTech developed RNAtag-seq as a method to reduce the cost, time and labour involved in RNA-seq library prep - allowing 96 samples in a plate to be pooled for single RNA-seq library prep.
The key to the method is the initial ligation of an RNA-barcode to each sample, then after pooling and clean-up a more standard RNA-seq prep is performed but on a pool of samples. Currently 32 samples are pooled for a Zymo column clean-up before RNA-seq library prep. Hopefully the protocol can be easily adapted to much larger pools using RNA SPRI or similar? Taking a 96 well plate of RNA samples, performing the RNA adapter ligation and a single library prep could allow a single lab tech to prepare 100's of samples in relatively short order.
The authors designed and tested a set of 96 RNA barcode adapters with a single E.coli samples to find those that showed the least ligation bias, and that gave the best read balance. However RNA ligase bias is well documented and some simple fixes to current protocols have been proposed (see previous post), I liked the random bases used in Tamas Dalmay's Silence 2012 paper and would have thought something similar would be an easier way to produce very large sets of barcodes.
To demonstrate the applicability of the data the authors presented two experiments. The first was a mouse differential gene expression analysis of tissue development, where they demonstrate the comparability to dUTP RNA-seq. In the second they identified a 67 gene signature of ciprofloxacin susceptibility in E. coli by profiling ciprofloxacin-susceptible and ciprofloxacin-resistant clinical isolates. I think it was a shame that they did not demonstrate how far the method can be pushed with a much larger experiment, but the current adapter design makes their ordering and testing expensive and time consuming. An 768 (8*96) sample RNAtag-seq experiment, with library prep in a single strip-tube would have been a very effective demonstration!
A note of thanks to the authors: I contacted the authors about their paper and had the kind of response you hope for but rarely get. My questions were answered and more besides. One thing that came out was the potential need to think carefully about which samples to pool. Since the only normalisation is the amount of total RNA input, significant differences in samples (rRNA:mRNA ratios, distribution of transcript expression and size etc) could result in very poor read balance between samples. However rather than spend too much time considering this I think I'd prefer to test this experimentally and decide what to do once I've got some data in front of me.
Livny also had a paper a couple of years ago looking at read-depth for bacterial RNA-seq. where they showed that just 5-10 million reads were sufficient to detect all but the lowest low expressed bacterial transcripts. Using one single-end 50bp HiSeq flowcell would allow 384 bacterial RNA-seq samples to be processed. This could conceivably be a single 384 well plate if enough barcodes are available.
RNAtag-seq explained: Most of us are making RNA-seq libaries using one of the strand-marking dUTP methods, where each library is taken through the steps of library prep in Eppendorf's or 96-well plates. Scientists at the Broad Institute and CalTech developed RNAtag-seq as a method to reduce the cost, time and labour involved in RNA-seq library prep - allowing 96 samples in a plate to be pooled for single RNA-seq library prep.
The key to the method is the initial ligation of an RNA-barcode to each sample, then after pooling and clean-up a more standard RNA-seq prep is performed but on a pool of samples. Currently 32 samples are pooled for a Zymo column clean-up before RNA-seq library prep. Hopefully the protocol can be easily adapted to much larger pools using RNA SPRI or similar? Taking a 96 well plate of RNA samples, performing the RNA adapter ligation and a single library prep could allow a single lab tech to prepare 100's of samples in relatively short order.
The authors designed and tested a set of 96 RNA barcode adapters with a single E.coli samples to find those that showed the least ligation bias, and that gave the best read balance. However RNA ligase bias is well documented and some simple fixes to current protocols have been proposed (see previous post), I liked the random bases used in Tamas Dalmay's Silence 2012 paper and would have thought something similar would be an easier way to produce very large sets of barcodes.
To demonstrate the applicability of the data the authors presented two experiments. The first was a mouse differential gene expression analysis of tissue development, where they demonstrate the comparability to dUTP RNA-seq. In the second they identified a 67 gene signature of ciprofloxacin susceptibility in E. coli by profiling ciprofloxacin-susceptible and ciprofloxacin-resistant clinical isolates. I think it was a shame that they did not demonstrate how far the method can be pushed with a much larger experiment, but the current adapter design makes their ordering and testing expensive and time consuming. An 768 (8*96) sample RNAtag-seq experiment, with library prep in a single strip-tube would have been a very effective demonstration!
 |
| RNAtag-seq strand assignment compares well to dUTP methods |
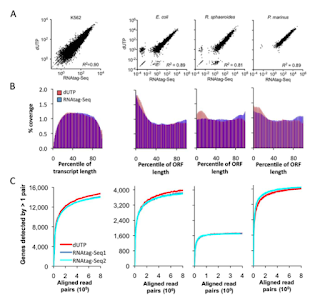 |
| RNAtag-seq compares well to dUTP methods on multiple metrics |
Livny also had a paper a couple of years ago looking at read-depth for bacterial RNA-seq. where they showed that just 5-10 million reads were sufficient to detect all but the lowest low expressed bacterial transcripts. Using one single-end 50bp HiSeq flowcell would allow 384 bacterial RNA-seq samples to be processed. This could conceivably be a single 384 well plate if enough barcodes are available.

I would be interested to know how one could use poly-a enrichment as opposed to ribosomal reduction. Did the authors comment on that to you?
ReplyDeleteIt will be no problem if you do polyA first and then library construction (pooling after barcoding and RT after pooling). But if you have hundreds of samples - easier to do rRNA depletion.
DeleteI'm missing something simple here. Why are there 2 sets of barcode oligos? One set of 32 DNA barcode oligos and a second set of 54 RNA barcode oligos?
ReplyDeleteDNA are cheaper per order, but RNA is much better (and cheaper per reaction).
Deleteis AR2 primer for cDNA synthesis same as the cDNA synthesis primer in ScriptSeq (Bacteria)? Here is the link for Scriptseq protocol. http://support.illumina.com/content/dam/illumina-support/documents/documentation/chemistry_documentation/samplepreps_truseq/scriptseq-complete/scriptseq-complete-kit-bacteria-library-prep-guide.pdf
ReplyDelete